Understanding the genetic bases of adaptation to soil water deficit in trees through the examination of water use efficiency and cavitation resistance: maritime pine as a case study
Abstract
This article provides a comprehensive view on the existing knowledge related to adaptation to soil water deficit in maritime pine, a conifer species widely planted in the southwestern Europe. It synthesizes discoveries made in ecophysiology, quantitative and population genetics as well as in genomics, combining several layers of information at the genotypic, phenotypic and environmental levels. Particular focus is given to two major traits: water-use efficiency (WUE) and cavitation resistance (CR). The former is related the maintenance of productivity during periods of lower soil water availability, whereas the latter is tightly linked to survival during severe drought. The development of high throughput phenotypic technologies have made it possible to estimate genetic and environmental variance components of these key traits, providing clues about their suitability for breeding and the evolutionary forces that have shaped their variability. Both CR and WUE were screened in different ecotypes as well as in the Aquitaine breeding population, the main genetic resource of the most advanced maritime pine breeding program in Europe.
While the unexpectedly low level of variation of CR within and between natural populations will most likely hamper its use in breeding application, for WUE the medium heritability, absence of unfavorable phenotypic and genetic correlations with diameter growth, as well as the high inter-site correlation and weak genotype-by-environment interaction indicates that artificial selection could be applied for this trait without unfavorable consequences for radial growth, at least within the Aquitaine provenance. On the other hand, recent advances in sequencing and genotyping technologies have contributed to reveal the genetic architecture (i.e. number, location and effect of quantitative trait loci) of these two traits. In combination with ultra-dense genetic linkage map and functional genomics approaches, these findings will contribute to identify positional and expressional candidate genes that should be validated by association genetics and eventually introduced in genomic prediction models to make such knowledge useful to improve tree breeding.
This review also opens up new research avenues and raises key questions on how to promote adaptation to the challenge of soil water deficit through genetic approaches in this species. If the results obtained so far in maritime pine can only applied to the tested genetic material, we however believe that the overall strategy presented here can be considered and cited as an example of integrative research to better understand the genetic bases of adaptation to soil water deficit in any forest tree species.
Introduction
Recent studies of global supply and demand for wood have confirmed that planted forests (see definition in FAO, 2012) are playing an increasing role in meeting wood and fiber needs. If we are to meet future demand, while concurrently reducing deforestation and disturbances in natural forests, we will therefore need to increase the sustainable productivity of existing planted areas and/or increase the rate of new plantations (by 2.4% according to recent estimates from the WWF and IIASA, 2012). Two complementary approaches can be used to improve the productivity of existing high-yield industrial plantations: i) the optimization of silvicultural practices, from plantation to harvest, taking into account carbon, water and nutrient cycling, and ii) the valorization of genetic resources through the restocking of local or non-local provenances or the use of exotic species for the breeding of improved varieties for deployment. Forest plantations have increased in area by over 110 million hectares over the last 25 years (MacDicken, 2015), and such plantations now account for 7% of the forested area worldwide (i.e., 290 M ha). As a result, about 50% of industrial roundwood is supplied by planted forests (Payn et al., 2015) and this proportion may increase considerably in the near future (Carle and Holmgren, 2008).
However, like all forest ecosystems, planted forests are subjected to the effect of climate change, with warmer temperatures and changes in precipitation patterns, the two main drivers of water status of forest stands. Drought and heat-related forest dieback is already reported on all continents, in both areas with water shortage and well-watered areas (Allen et al., 2010; 2015). Hydraulic failure (i.e., desiccation due to a failure of water transport) and carbon starvation (when carbon acquisition and the mobilization of carbon stores fail to meet metabolic maintenance needs) have been recognized to play a role in this widespread phenomenon (Hartmann, 2015). It was also recently demonstrated that many forests and woodlands are facing an increasing risk of climate-induced dieback (Choat et al., 2012). Thus, climate change and associated natural disturbances are likely to become dominant stressors, particularly if temperatures increase substantially (Settele et al., 2014). As a consequence, net primary production from planted forests would be expected to decrease in regions in which increases in water deficit during the growing period are projected, such as the south-western part of Europe (Loustau et al., 2005). Similar changes will also result from severe heat stress, as observed in Western Europe in the summer of 2003 (Ciais et al., 2005; Breda et al., 2006).
The choice of planting stock during restocking and the management of existing plantations are two important elements in adaptation to climate change (Lindner et al., 2010). Decisions concerning the suitability of a particular species/provenance/variety in altered climatic conditions should have a sound scientific basis. There is therefore an urgent need to address a number of questions: Which tree functional traits are most relevant for adaptation to drought? How can we measure these traits with high-throughput techniques compatible with the sample sizes required for the screening of genetic resources, the estimation of classical genetic parameters and the selection of improved genotypes? To what extent are these traits determined genetically and environmentally? What do we know about the genetic architecture of these traits, the nature and regulation of the genes/gene networks involved? Moreover, if we assume that the traits (and underlying genes) likely to be under selection in our changing environment have probably been under natural selection in the past (i.e. during warm periods following the last glacial optimum), then understanding how species responded to past climate change could help us to forecast the response of species in the future. Improvements to our understanding of the evolutionary trajectories of native forest tree populations should therefore facilitate: i) the selection of genetic resources already adapted to future environmental conditions, and ii) the adoption of specific measures, such as the assisted migration of climate-adapted genotypes (Richardson et al., 2009). We therefore need to find answers to the questions raised above, to enable us to establish resilient forest ecosystems that can continue to supply a broad range of services.
In this review article, we tackle two major challenges concerning adaptation to water deficit in these long-lived organisms: the maintenance of productivity with less water availability and survival during severe drought or heatwaves. We focus in particular on the characterization of water-use efficiency (the ratio between accumulated biomass and transpired water) and cavitation resistance (the ability to conduct water though the xylem even during drought events). We consider three types of information, relating to the genotype, phenotype and interaction with environment. Our knowledge of genotypes and phenotypes is continually being increased by improvements on genotyping and phenotyping technologies. We focus on observations and experimental results (listed in supplementary file 1) for a single species of ecological and economic importance in the southwestern Europe: maritime pine (Pinus pinaster Ait.). This species is considered to be a drought-avoiding species with sensitive stomata, displaying rapid osmotic adjustment in response to water deficit (Granier and Loustau, 1994; Picon et al., 1996). It also has a large ecological amplitude in its natural range, with contrasting ecotypes, relating to water availability in particular (Nguyen and Lamant, 1989; Fernández et al., 1999; Fernández et al., 2000). This species extends over 4M ha in its natural range and has been subject to an active breeding program in France since the early 1960s (Illy, 1966). Despite this focus on one species, we believe that this research approach will prove valid for other biological systems, therefore providing a generic framework in which to study the genetic bases of adaptation to water deficit in other forest tree species.
The three building blocks for studying the genetic basis of adaptation to water deficit
Environmental data
Several levels of environmental variability and its impact on the phenotype can be considered. In classical quantitative genetics studies, environmental variations are often taken into account and minimized through the use of specific experimental designs with blocks (for mesoenvironmental variations) or spatial correction techniques (for microenvironmental variations). For example, in the Papadakis method (Papadakis, 1984), the residual value of each neighboring individual included in a grid of appropriate dimensions is used to adjust individual values for microenvironmental effects. These methods do not require environmental measurements per se. Conversely, the consideration of macro-environmental variability is important in studies of reaction norms (range of phenotypes produced by a particular genotype in different environmental conditions) and in cases in which its variance component must be estimated. This is for example the case for the modeling of phenotypic data over different environments, the explicit interpretation of genotype-by-environment (Denis, 1988), the analysis of quantitative trait loci (QTL)-by-environment interactions or inter-site genetic correlations. In population genetics studies, it is essential to characterize the habitat in which populations have evolved over generations of natural selection, to infer: i) adaptive phenotypic variation, from the correlation between trait variation (e.g. measured in common garden experiments) and environmental variation due to differences at the populations of origin, or ii) adaptive genomic variation, from the correlation between environmental variables at the populations of origin and genetic polymorphisms. In such “environmental” associations (Coop et al., 2010; Frichot et al., 2013), it is implicitly assumed that natural selection had enough time to create a functional relationship between allele frequencies and environmental variables.
The environmental attributes that should be considered include: i) coarse-grained abiotic features, such as classic pedoclimatic variables (e.g. air temperature, precipitation, evapotranspiration, irradiation, soil type, pH, organic matter concentration, soil moisture, soil water storage capacity, soil mineral content), including indices obtained by combining several variables relating to drought (Vicente-Serrano et al., 2012; Pedro-Monzonís et al., 2015), ii) topographic information (e.g. elevation, gradient, aspect, form of the slope), and iii) more complex biotic features characterizing ecological niches. Habitat description can be improved by using reverse models of community ecology to predict local environmental conditions from the abundance and assemblies of the plant community. Ecologists have been using such approaches for years, in the form of Ellenberg indices for particular species, but this method has several limitations in that it produces mean class values and the quality of the relationship is specific to each ecosystem (e.g. (Dzwonko, 2001; Wamelink et al., 2002; Szymura et al., 2014). Similarly, the description of microbiome biodiversity through metagenomic approaches is also becoming increasingly powerful for characterizing the environmental setting in which populations are sampled (Tringe and Rubin, 2005).
Such environmental data are frequently not collected in field tests or in the populations of origin, and, when available, they are often limited in scope or resolution. In the genetic studies considered in this review, environmental data were either inferred from vegetation data, as for the Aquitaine populations in the study by Marguerit et al. (2014), or based on climatic data (e.g. monthly mean, highest, and lowest temperatures and mean precipitation) obtained from global (such as Worldclim, Hijmans et al., 2005) and/or local monitoring systems, as described by Lamy et al. (2014) and Jaramillo-Correa et al. (2015a). This lack of precise environmental characterization contrasts strongly with the ever-increasing rate at which we are able to collect genomic and phenotypic information (see below), making it difficult to make evolutionary inferences from the analysis of genotype-environment and phenotype-environment datasets. The characterization of relevant ecological pressures acting on individual phenotypes is therefore a key issue that must be addressed.
Genotyping technologies
In recent years, groundbreaking technologies have been developed for the characterization of DNA sequence variation within a population (Ekblom and Galindo, 2011). These tools have greatly accelerated the characterization of forest tree diversity and helped to broaden the genetic base of breeding populations for improving adaptation to climate change. There are a number of classes of genetic variation. The smallest observable unit of DNA polymorphism and the type of variant most frequently found throughout the genome, involves differences of a single nucleotide (A, T, C or G) between individuals (single-nucleotide polymorphism or SNPs). Highly multiplexed SNP genotyping assays can be used to characterize the genotypes (heterozygous or homozygous) of hundreds or thousands of individuals at thousands of loci in parallel, and then to estimate allele frequency in a given population. This metric is frequently used for population genetic inferences. For maritime pine, catalogs containing hundreds or thousands of SNPs have been developed by amplicon sequencing, targeting candidate genes (Pot et al., 2005; Eveno et al., 2008; Lepoittevin et al., 2010; Grivet et al., 2011; Grivet et al., 2013) or by mining expressed sequence tags (Chancerel et al., 2011; Chancerel et al., 2013; Canales et al., 2014). For this species, Illumina bead array technology (Illumina, San Diego, CA, USA) is the principal tool used (Plomion et al., 2016). This technology requires only information about the flanking sequence of a known SNP to design an oligonucleotide probe (attached to a bead) specific for this sequence binding just upstream from the targeted SNP. The probe hybridizes to the 50 bases flanking the SNP in a genomic DNA sample and base extension with fluorescently labeled bases is then used to investigate the SNP (each base has a different color). A scanner is used to image the array, on which hundreds of thousands of assays can be spotted (Supplementary file 2).
High-throughput genotyping is still expensive but is rapidly becoming more affordable, with the development of genotyping-by-sequencing technologies (Deschamps et al., 2012), making it possible to perform genetic studies with very few assumptions about the genes underlying phenotypic variation, and unbiased genome-wide searches for genes of interest (e.g. in poplar by Evans et al. (2014)). Exome capture and direct sequencing (Gnirke et al., 2009) techniques are currently being developed for maritime pine. This technology represents a great step forward in terms of the numbers of markers available, but the direct sequencing of individuals of interest associated with this approach may also attenuate one of the two main problems associated with current genotyping assays: i) ascertainment bias due to the small number of individuals from which SNPs were obtained (ascertainment width) and ii) the stringent criteria for minimum allele frequency and read coverage used to retain the SNPs (ascertainment depth).
Which traits are relevant?
Phenotyping techniques have also received a boost from technological developments facilitating the measurement of phenotypes of interest. As mentioned above, the key challenges are identifying genotypes with a higher growth efficiency under moderate drought conditions and/or genotypes more resilient to extreme drought events. The mechanisms underlying these two processes are different, and the specific traits involved therefore needed to be identified.
Water use efficiency (WUE)
Unlike herbaceous plants, perennial and woody plants, such as forest trees, cannot completely escape drought periods. However, they can adjust their annual growth cycle to annually reoccurring soil water deficit, by modifying the timing of budburst and leaf development and through drought-induced leaf shedding. Therefore, the drought resistance of trees in natural forests is mostly a question of survival and competition within an ecosystem. Tree growth performance may also be related to the ability of maintaining growth during periods of water shortage. Plants can develop different strategies to avoid or tolerate soil water deficit. Some of the most rapid acclimation responses of trees to soil water deficit include decrease in diameter and height growth, and stomatal closure, followed by morphological adjustments, such as an increased root growth (Bogeat-Triboulot et al., 2007). The exploration of available soil moisture is particularly important for Mediterranean plants, such as maritime pine. Stomatal closure at the onset of drought enables the plant to maintain hydraulic integrity. However, stomatal closure can also decrease the leaf internal CO2 availability, thereby decreasing either growth or carbon reserves. The delay of stomatal closure with increasing soil water deficit varies among Mediterranean tree species (Galmés et al., 2007). This balance between growth and water use during a drought stress can be characterized by calculating the ratio of net CO2 assimilation rate (A: as a proxy for production) to stomatal conductance for water vapor (gs; as a proxy for transpiration), which is called the intrinsic water use efficiency Wi (Meinzer et al., 1991). The Wi of maritime pine seedlings increases considerably over a soil drying cycle, as does the variation within a full-sib family (Marguerit et al., 2014 ; Figure 1). Wi can also be estimated by stable carbon isotope analysis on organic material from the plant (δ13C) based on the model linking δ13C with Wi developed by Farquhar et al. (1989). Depending on plant component used, the carbon isotope composition of the plant material may integrate Wi over time, resulting in a more stable measure of plant specific values. This also makes the large-scale screening of genetic material possible, through the harvesting of leaf or wood material. Wi estimates water-use efficiency at the leaf level, whereas in tree breeding, increases in biomass production are an important goal. It is therefore important to consider the improvement in Wi for a tree in terms of effects on plant growth capacities (Dvorak, 2012).
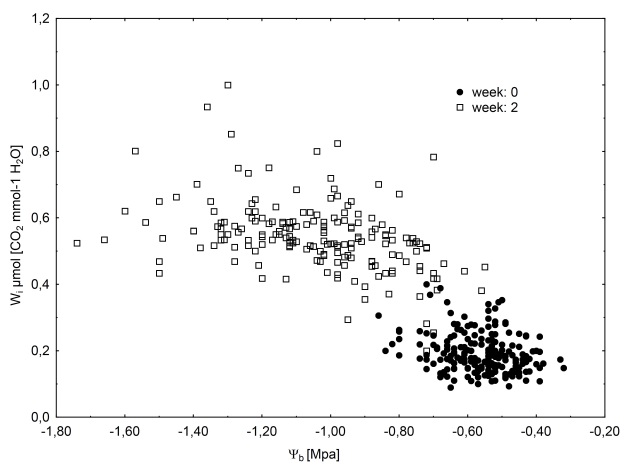
Figure 1. Relationship between predawn water potential (as an estimator of soil water deficit) and intrinsic water-use efficiency (Wi = A/gs) for a two-week drying cycle. The data shown were obtained from a maritime pine family described by Marguerit et al. (2014): measurements before stress application (well-watered two-year-old seedlings) in black circles; measurements after two weeks of soil drying in white squares.
Cavitation resistance (CR)
One of the most important mechanisms leading to plant death during drought is hydraulic failure of the plant water transport system due to the development of excessive tensions in the xylem under severe drought stress conditions (Delzon and Cochard, 2014). Anderegg et al. (2015) recently showed that tree mortality can be predicted exclusively from the loss of vascular transport capacity. This hydraulic failure results from catastrophic increases in xylem cavitation, the change from liquid to water vapor caused by the high tensions required for water transport from the soil to the leaves (Tyree and Zimmermann, 2002). As the percentage of cavitated xylem conduits increases, the hydraulic conductivity of the xylem decreases, until the flow of water stops, causing desiccation of the tree tissues. This desiccation leads to the death of affected leaves and branches and, in particularly severe cases, may result in the death of the tree. Brodribb and Cochard (2009) and Brodribb et al. (2010) assessed the link between cavitation and plant death for seven southern-hemisphere conifer species with different levels of drought tolerance. A strong link between the xylem pressure inducing a 50% loss of stem conductivity (called P50) and the lethal water potential was observed (Figure 2).
Species growing in dry environments are more resistant to drought-induced cavitation (more negative P50 values; Maherali et al., 2004; Cochard et al., 2008; Choat et al., 2012). Resistance to cavitation is therefore a critically important trait for evaluating the ability of trees to survive and recover from drought periods and, therefore, for predicting future drought-induced forest decline. Cavitation resistance is known to be linked to the anatomy of xylem conduit walls, and, more specifically, to that of the pit walls. Xylem conduits are composed of dead cells, so this trait is determined at the time at which the conduits are formed and remains an intrinsic propriety of the conduits as long as they remain functional.
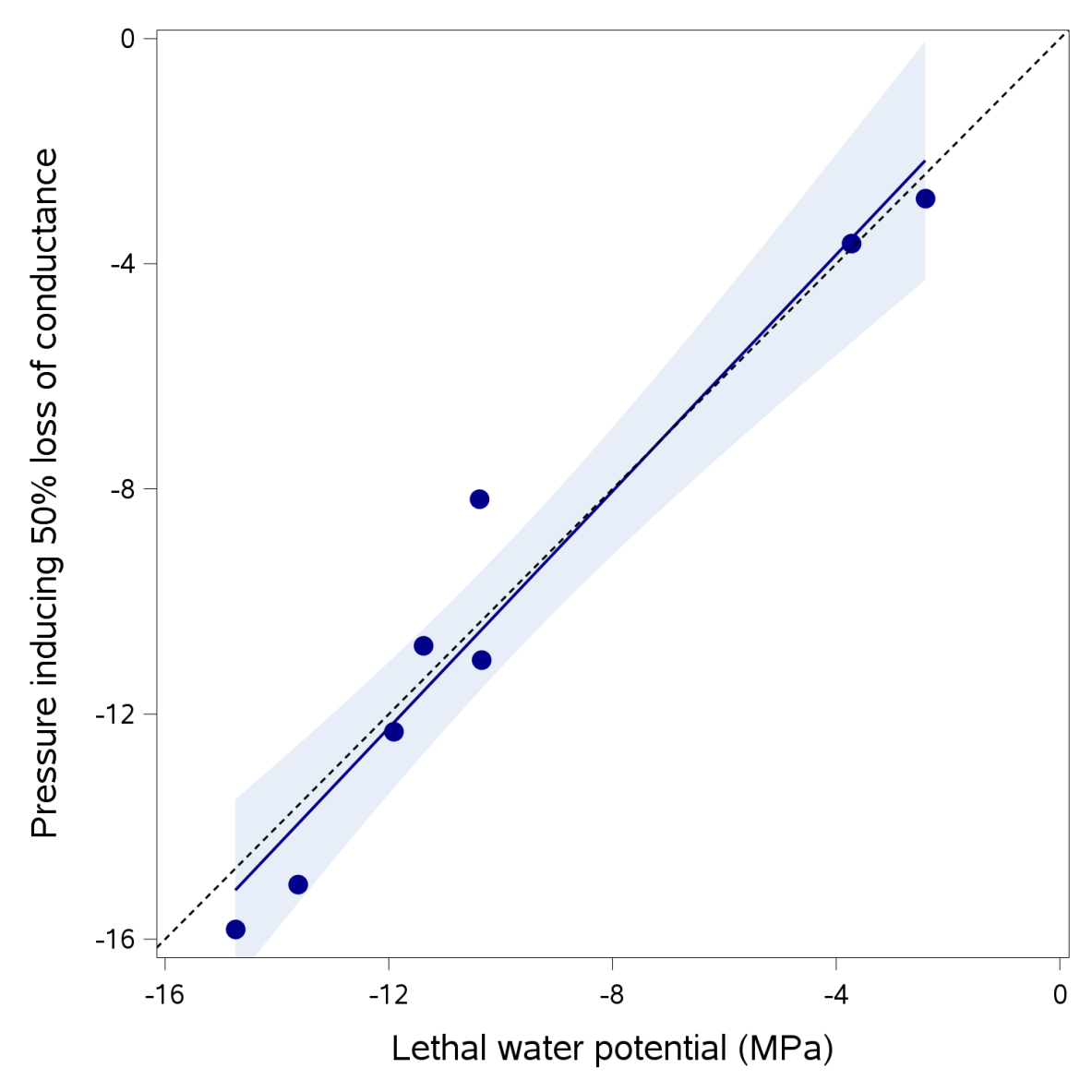
Figure 2. Illustration of the highly significant 1:1 relationship between P50 (xylem pressure inducing 50% loss of hydraulic conductance, measured with the Cavitron technique) and lethal water potential, as determined by gas exchange recovery kinetics; r2 = 0.95). Each dot corresponds to a different conifer species. Data retrieved from Brodribb and Cochard (2009) and Brodribb et al. (2010). The correlation coefficient is given for the regression line. The dashed line corresponds to the 1:1 relationship.
Ecophysiologists generally characterize xylem resistance to cavitation by using vulnerability curves (VCs) calculated from the relative flow rates measured on xylem segments. VCs show how the percentage cavitation in a xylem tissue varies with xylem pressure. Several laboratory-based techniques have been developed for the plotting of VCs (see Cochard et al., 2013 for a review), but most are time-consuming and of low sensitivity, making it impossible to screen and compare populations or genotypes. Over the last decade, a new technique based on the centrifugation of a xylem segment (in a Cavitron) has been developed, making it possible to generate VCs rapidly (Cochard et al., 2005). Five years after the development of this technique, high-throughput phenotyping facilities for cavitation resistance were established in France (Clermont-Ferrand and Bordeaux, http://sylvain-delzon.com/caviplace/, Figure 3), for assessing the relationship between phenotype and genotype.
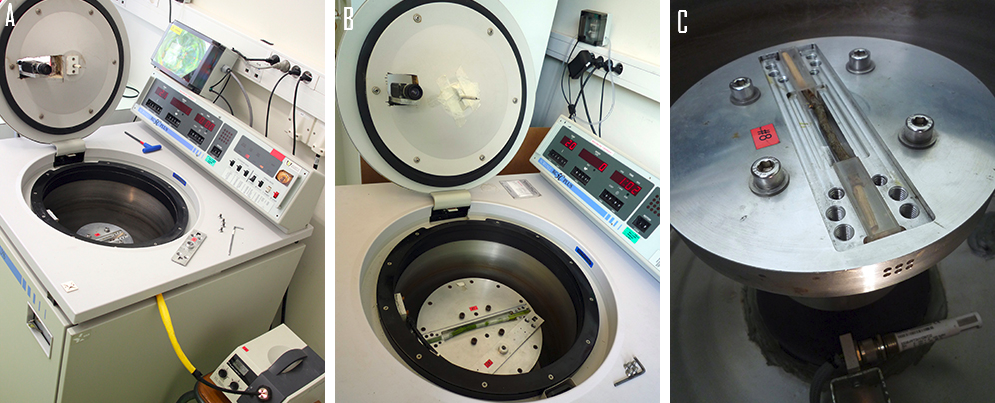

Figure 3. Standard Cavitron showing the centrifuge and the light source on the left (A) and the center and the rotor with the branches (B) and the two water reservoirs on the right (C). The flow centrifuge technique (also called the Cavitron) uses a centrifugal force to create a negative xylem pressure that induces embolism in the sample. Samples are placed in a custom-built rotor with both sample ends in reservoirs filled with water. In order to generate a pressure gradient across the sample and thus induce water flow through the sample, small holes are made in the reservoirs at two distances from the rotation axis (Cochard et al., 2005).
In total, six “standard” Cavitrons (standard rotor = 26 cm in diameter) are used to study and quantify intraspecific variability in tracheid-bearing species. The VC for a sample can typically be obtained in less than half an hour with this technique, making it possible to screen large number of genotypes for cavitation resistance. Just to provide some idea, more than 1,500 VCs have been produced for maritime pine (P. pinaster) over the last three years, from branches sampled from common gardens (provenance and progeny tests) and from natural populations in situ (Lamy et al., 2014). It is worth noticing that the variability of P50 within a tree is very low, rendering the assessment of cavitation resistance highly robust. Recent results show indeed similar resistance to cavitation of the xylem in branches and leaves in Pinus pinaster (Bouche et al., 2015), and in any case a very strong relationship between branch P50 and leaf P50 exists in conifers (Delzon, pers. comm.). In addition, a recent study reported very low intra-tree (branch, trunk and root) variability for P50 in several conifer species including maritime pine (Bouche et al., 2016).
The Cavitron technique is a considerable improvement over previous more laborious techniques, but it remains time-consuming and not widely accessible, this technique being used in only in a handful of highly specialized laboratories around the world. High-throughput phenotyping of cavitation resistance will probably be dependent on the development of groundbreaking methods very different from those currently used. Preliminary tests have suggested that near-infrared spectra obtained from ground wood from different species can be used to predict P50 values. However, this approach remains to be validated for the screening of genotypes within a species.
WUE and CR may thus be considered promising traits for assessing the responses of trees to mild drought and their survival/resilience in the face of intense water deficit, respectively. Selection for higher resilience may come at the expense of a lower growth efficiency under conditions of moderate stress. This hypothesis may hold true at the interspecific level (Hacke et al., 2001), but the results of the very few studies addressing this key issue at the intraspecific level have been less conclusive. For instance, in Populus deltoides X nigra hybrids, Fichot et al. (2010) found no correlation between δ13C and P50. By contrast, Lamy et al. (2011) found a significant correlation between these two traits across Pinus pinaster provenances, with the more cavitation-resistant genotypes tending to have lower water use efficiency. This correlation clearly requires further investigation, but, as the link between these two traits is mechanistically weak, the chances of identifying 'elite' genotypes combining high performance for both traits are high.
Other traits shedding new light on the genetics of adaptation to water deficit
The following three examples highlight current efforts to improve the functional characterization of the adaptation of forest trees to water deficit, with the aim of developing new methods or using already tested methods compatible with the large sample sizes required for genetic studies.
A first option will be to identify one or several anatomical traits of the xylem linked to CR at the intraspecific level. Within coniferous taxa, the tremendous variability of CR is strongly associated with the bordered pit structure in tracheids. Indeed, recent studies (Delzon et al., 2010; Pittermann et al., 2010; Bouche et al., 2014) have shown that the size of the torus relative to the pit aperture (also called torus overlap) is closely related to cavitation resistance. This observation has led to calls for further studies assessing whether there is also a relationship between P50 and pit properties between genotypes within species. If so, torus-aperture overlap in conifer pits may be a good proxy for assessing genetic variation in drought tolerance. However, observations of pit anatomy require the use of a scanning electron microscope, and this approach remains time-consuming. However, it has the advantage that it can be applied to small wood samples.
A second approach involves the assessment of foliar traits for the identification of individuals capable of continuing to fix carbon or of recovering this ability during and after stress. Fluorescence measurements are an interesting alternative to gas exchange measurements for the quantification of photosynthetic activity (Hunault et al., 2015). Phenotyping can then be performed with classic Fv/Fm and electron transport rate measurements or by measuring the rapid transient response to flash lights (Munns et al., 2010; Furbank and Tester, 2011). These techniques have been implemented in crops (e.g. wheat, rice, pea, barley), to compare the photosynthetic functioning of different varieties or genotypes in response to abiotic stresses (drought, heat). They can easily be implemented in the field and on large sample sizes. Classical fluorescence indices have been used to characterize G x E interactions in several forest trees and have revealed differences between pine provenances (Michelozzi et al., 2011; Reinhardt et al., 2011). Moreover, fluorescence measurements and xanthophyll cycle pigment concentrations can also be used to estimate non-photochemical quenching (NPQ). These methods can be used to quantify inter- and intra-specific variations of the heat dissipation energy that helps to protect photosystems during drought in five Quercus species (Ramírez-Valiente et al., 2015). Studies using these methods have demonstrated the existence of a link between pigment physiology and stress tolerance at the intra-specific level. These promising methods are now being applied to maritime pine.
A third breakthrough in investigations of the genetic and environmental determinism of forest tree adaptation to water deficit is the development of measurements for analysing the dynamics of plant behaviour in response to environmental conditions. High-resolution stem diameter variation (SDV) is an effective tool for studies of the fundamental mechanisms underlying whole plant functioning and growth, including plant water relations, plant hydraulics, plant carbon relations, plant nutrition, freezing effects, plant phenology and dendroclimatology (reviewed by de Swaef et al., 2015). SDV patterns are commonly characterized by fixing point or band dendrometers on the trunk to study both tree water status and radial growth (Steppe et al., 2008; Zweifel et al., 2010; King et al., 2013). SDV patterns and their association with wood density fluctuations have been also analyzed in different environmental conditions (Drew et al., 2009; 2011). Using band dendrometers and weekly monitoring of intra-annual density fluctuations in maritime pine, Vieira et al. (2015) showed that cambial activity adjusted to environmental conditions, with the resumption of woody cell production after the summer drought, as a function of precipitation events. Tools for measuring SDV in a large number of genotypes, with automated data acquisition systems, are providing new perspectives for deciphering genotype-by-environment interactions in forest tree genetics. Indeed, characterizing the dynamics of SDV within populations should make it possible to identify the key genetic components of this plastic response. This approach is being applied to maritime pine, with 250 trees (from two full-sib families) equipped with automated microdendrometers measuring daily fluctuations in stem diameter (Lagraulet, 2015). Investigations of the genetic architecture of such daily traits should, in the long term (over several years), make it possible to determine the dynamics of the genetic control of maritime pine behaviour as a function of environmental conditions.
Variance components of the phenotypes and opportunities for breeding
The genetic basis of Wi has been investigated in several provenances of maritime pine, by evaluating δ13C levels in wood and needles. Brendel et al. (2002) obtained the first estimate of additive genetic variability for δ13C in a half-diallel set-up with 12 genotypes from the Landes provenance. They measured δ13C levels in cellulose extracted from four successive annual wood rings (11 to 14 years old) with an IRMS (isotope ratio mass spectrometer). The heritability (h²) of δ13C was estimated at 0.17 and was very similar to that of ring width (h²=0.19). A more recent study based on a more genetically diverse sample from the same population of the Landes provenance (i.e. 50 half-sib families with a low level of relatedness) yielded a higher estimate of heritability: Marguerit et al. (2014) found significant additive genetic variability for this population, with an h² of 0.29 ± 0.07. The methods used were similar to those used in the previous study: wood cores from 10 to 13 trees per family (1,901 trees sampled) and three successive annual rings (10 to 12 years old). The level of heritability was similar to or higher than that for growth-related traits in the same population (h² = 0.25 ± 0.07). This second estimate was based on the largest data set reported to date for a forest tree species. Another study based on six different provenances (including the Landes provenance) from France, Spain and Morocco gave similar results (Lamy et al., 2011). In this study, δ13C levels were assessed in the needles of six-year-old trees and a heritability of 0.21 was obtained for δ13C. This value is between those obtained in the other three studies.
For maritime pine breeding, the absence of an unfavorable correlation with diameter growth found within in the Aquitaine germplasm (Marguerit et al., 2014) was a key finding, suggesting that past selection for growth has not decrease the WUE of the trees and that artificial selection for WUE could be applied without unfavorable consequences for radial growth potential, at least within this provenance. The inter-site correlation for δ13C was high in this study (0.63-0.91), with a significant but weak genotype-by-environment interaction, adding another asset for breeding. However, even though a favorable level of heritability was detected, the relatively low level of genetic variability may limit the expected genetic gain. Marguerit et al. (2014) estimated that the mean δ13C of the breeding population (-26.1‰) would shift 0.4‰ if the best 1% of plants were selected. Genetic variability could be introduced into the breeding population from other provenances, as large differences in δ13C and WUE have been shown among populations (Guehl et al. 1995). More specifically, Marguerit et al. (2014) showed that a recessive allele from the Corsican-Restonican background could improve WUE. A new experimental design specifically established with Corsican populations (30 populations*30 half-sib families/population*50 offspring/family) will be used in the near future, to evaluate the genetic variability for WUE in the Corsican provenance.
As described above, only two studies have investigated the genetic component of cavitation resistance in maritime pine (Lamy et al., 2011; 2014). These studies were based on maritime pine provenances established in common gardens. The heritability of P50 was found to be high: 0.44 in the study by Lamy et al. (2011) and 0.51 to 0.61 in the study by Lamy et al. (2014), contrasting with the low to moderate heritability found for growth and WUE in the same studies. However, the most striking result was the extremely low level of variation within and between populations, decreasing the chances of genetic improvement by breeding.
Linking phenotype and environment: how do phenotypes fit their local ecological niches?
Forest trees are a classic example of species with strong phenotypic responses to environment, for which phenotypic clines in response to geographic or environmental variables have been reported for several traits (e.g. see Wahid et al. (2006) for needle traits in maritime pine). The plastic component of this response is not negligible (and can also be adaptive), but both classical and next-generation genecological studies have shown that local adaptation frequently underlies genetic variation across environments (Savolainen et al., 2007; Morgenstern, 2011). Local adaptation in forest trees can result in genetic clines reflecting a balance between gene flow and natural selection (Barton, 1999). Furthermore, in species that have survived periods of glaciation in isolated multiple glacial refugia, such as maritime pine, ecotypic differentiation is also common. Alberto et al. (2013) recently reviewed the evidence for genetic clines in forest trees, focusing on phenological traits. Standard proof of genetic-based adaptation to the environment is generally obtained by demonstrating a correlation between the population mean phenotypic or genetic values obtained in common gardens and environmental variables from the source population. However, only multi-site experiments or reciprocal transplants constitute ideal designs to disentangle local adaptation (i.e. G: genetically based phenotypic divergence) from phenotypic plasticity (E) and its genetic component (G×E).
Water use efficiency and related traits have been studied for several maritime pine provenances. French Atlantic provenances (Landes, Medoc) have been compared with Corsican (Porto Vecchio) and North African (Tamjout, Tamrabta, Tabarka) provenances. Guehl et al. (1995) showed that trees from the Tamjout provenance had a lower WUE, both for whole plants (transpiration efficiency was estimated directly from whole-plant water loss and biomass accumulation) and for leaves (Wi and δ13C), than trees from the Medoc and Porto Vecchio populations, even though the latter two provenances were adapted to lower levels of precipitation and higher temperatures at their sites of origin. Similarly, Nguyen-Queyrens et al. (1998) found a lower mean WUE over 16 years for cellulose extracted from tree rings, for two genotypes from Tamjout compared to two genotypes from the Landes provenance. These differences were significant under both control and drought conditions, except for Wi, which nevertheless displayed a similar trend between the selected genotypes of the two provenances. However, the trees from Tamjout displayed a greater acclimation of leaf WUE to drought than those from the other two provenances, whereas acclimation was less strong at the whole-plant level (TE). This suggests that trees from the Tamjout provenance display a strong stomatal reaction between control and drought conditions. Guehl et al. (1995) also showed that trees from Tamjout grew less strongly than trees from the other two provenances, but that they invested more biomass in their root systems, under both drought and control conditions. Nguyen (1989) showed that roots from Tamjout seedlings grew faster in osmotic stress conditions and displayed greater osmotic adjustment than roots of seedlings from the Landes provenance. In leaves, osmotic adjustment to decreasing relative water content in the needles was found to be strongly correlated with mean annual rainfall at the site of origin (Nguyen and Bouchet-Lannat, 2003). In this study, the Tamjout provenance had the greatest capacity for osmotic adjustment. Trees from the Tamjout provenance therefore seem to invest more carbon in soil water exploration and the maintenance of cell turgor through osmotic adjustment, making it possible to open the stomata more strongly when water is available and to close them more strongly when water resources are limited. This results in a non-intuitive lower water use efficiency for this dry-site provenance, as also reported for the Moroccan Tamrabta provenance by Aranda et al. (2010) and Lamy et al. (2011). Aranda and coworkers also showed that within-population variation was lowest for this population, without the significant increase in WUE in the presence of soil water deficit, as reported for other provenances. Similarly, Tognetti et al. (2000) showed a strong provenance-by-drought interaction for WUE for different provenances from Portugal, Italy, Sardinia and Corsica. These findings suggest that there is considerable diversity in the drought-response strategies of maritime pine provenances. The data acquired to date suggest that some maritime pine populations may have adapted to water deficit by increasing WUE (e.g. Iberian populations, where mesic and xeric provenances have WUE values consistent with the climate of the source population; Corcuera et al., 2012), whereas others, such as those from North African provenances, may have developed mechanisms of drought avoidance (e.g. by investing more in the root system; Guehl et al., 1995; Aranda et al., 2010), as well as of drought tolerance through enhanced osmotic adjustment ability (Nguyen-Queyrens and Bouchet-Lannat, 2003).
Unlike WUE, CR (as evaluated by P50) seems to vary little between populations and displays no significant overall association with climate, at either the phenotypic or genetic level, possibly due to uniform selection or canalization (Lamy et al., 2014). Nonetheless, Lamy et al. (2011) found the Moroccan provenance to have the highest resistance to cavitation in a study of six populations sampled along an environmental gradient. Evaluations of CR in common gardens from wet and dry sites showed that phenotypic plasticity in P50, although significant, was relatively weak and not always correlated with the climate of the provenance of origin. One study found CR to be higher for the wet site than for the dry site (Lamy et al., 2014), whereas another reported the opposite pattern (Corcuera et al., 2011). The assessment of plant plasticity for resistance to cavitation is challenging, as vulnerability curves cannot be obtained directly for water-stressed plants due to native embolism. The use of a proxy, such as xylem anatomical traits, might therefore be a good alternative in this case.
Linking genotype and phenotype: identification of loci/genomic regions underlying phenotypic variation
A more mechanistic approach to studying the genetic basis of adaptation involves the partitioning of phenotypic variance into genetic and non-genetic components, through the linking of genotypes and phenotypes. There are several possible pitfalls in this approach, because phenotypic variation of most adaptive traits does not follow a simple inheritance pattern (Mauricio, 2001). Quantitative adaptive traits are controlled by the expression and interaction of several genes modulated by environmental conditions to different degrees, depending on the trait studied (Mauricio, 2001; Visscher et al., 2008). Complex adaptive traits can be dissected through two approaches, as described in Supplementary file 3: linkage disequilibrium mapping (also known as association mapping) and QTL mapping. Both approaches aim to find statistically significant associations between a genetic marker and a phenotype. Association mapping involves the use of population allele frequencies to determine the relationship between observed phenotypes and genotyped genes, whereas QTL mapping is based on the segregation of alleles in pedigrees of known structure.
QTL mapping has been used in forest trees for the last 20 years (see Jaramillo-Correa et al., 2015a for a review on adaptive traits). With the recent advent of high-throughput genotyping technologies, ultra-dense gene-based linkage maps have been constructed for major tree species (e.g. oak: Bodénès et al., 2016; pine: de Miguel et al., 2015; spruce: Pavy et al., 2012; eucalypts: Bartholomé et al., 2015a), leading to the identification of positional candidate genes with map locations coinciding with trait-QTLs (e.g. for carbon isotope discrimination in eucalyptus (Bartholomé et al., 2015b) and poplar (Monclus et al., 2012)). However, the large confidence intervals surrounding the QTL peak have led to studies of the genetic control of adaptive traits shifting to association mapping, making use of the large number of recombination events occurring over many generations in natural populations to narrow down the positions of loci controlling traits of interest (Jaramillo-Correa et al., 2015a). This approach is based on the assumption that only tightly linked loci display statistically significant association with the phenotypic trait after many generations and random mating. The principal drawback is that large numbers of genotyped molecular markers are required for fine analyses at the genome level (Neale and Savolainen, 2004), because conifers have enormous genomes (Mackay et al., 2012; Nystedt et al., 2013), with rapidly decaying linkage disequilibrium (Isik et al., 2015). In addition, phenotyping capacity may also become limited because of the large sample size for the detection of QTLs with a weak effect. These two features have made association mapping particularly laborious for these species.
The genetic control of adaptive traits in maritime pine has been studied through these two approaches. First QTL studies have focused on important productive traits, such as growth and wood quality (Plomion et al., 1996a; Chagné et al., 2003; Pot et al., 2006), or plant defense-related traits, such as the production of terpenes (Plomion et al., 1996b). Association studies in maritime pine have also been carried out with candidate genes, and two SNPs associated with growth and wood chemistry (Lepoittevin et al., 2012) were identified, together with 17 SNPs associated with serotiny (Budde et al., 2014). Only in recent years has it become possible to analyze functional traits related to drought tolerance, such as WUE, through QTL analysis (Brendel et al., 2002; de Miguel et al., 2014; Marguerit et al., 2014). These studies dissected the genetic architecture of WUE estimated through gas exchange (Wi) and δ13C. Vegetative propagation is challenging in maritime pine and most of these studies were based on single-tree estimates of WUE, except in the study by de Miguel et al. (2014). In general, few QTLs for WUE have been detected, with those found having only moderate effects. The only exception was a major QTL for δ13C detected in linkage group #12 by Marguerit et al. (2014) and accounting for 67% of the observed phenotypic variance. The use of a unique F2 pedigree with contrasting alleles from different provenances (Landes vs. Corsica) certainly contributed to the detection of this major gene. This major QTL colocalized with a QTL for Wi estimated by direct gas exchange measurements. This result is interesting, because the colocalization of QTLs for different traits may provide evidence of a functional relationship between those traits. The use of markers common to the genetic maps used in the three previously reported studies made it possible to compare the locations of QTLs for WUE in different genetic backgrounds, under different environmental conditions and at different developmental stages. Several QTLs co-localized for at least two independent studies in linkage groups #5, #6, #8 and #12 (Brendel et al., 2002; de Miguel et al., 2014; Marguerit et al., 2014). Figure 4 shows the map locations of all the QTLs for WUE detected to date, based on the P. pinaster composite linkage map from de Miguel et al. (2015) as a common framework for projecting these QTLs.
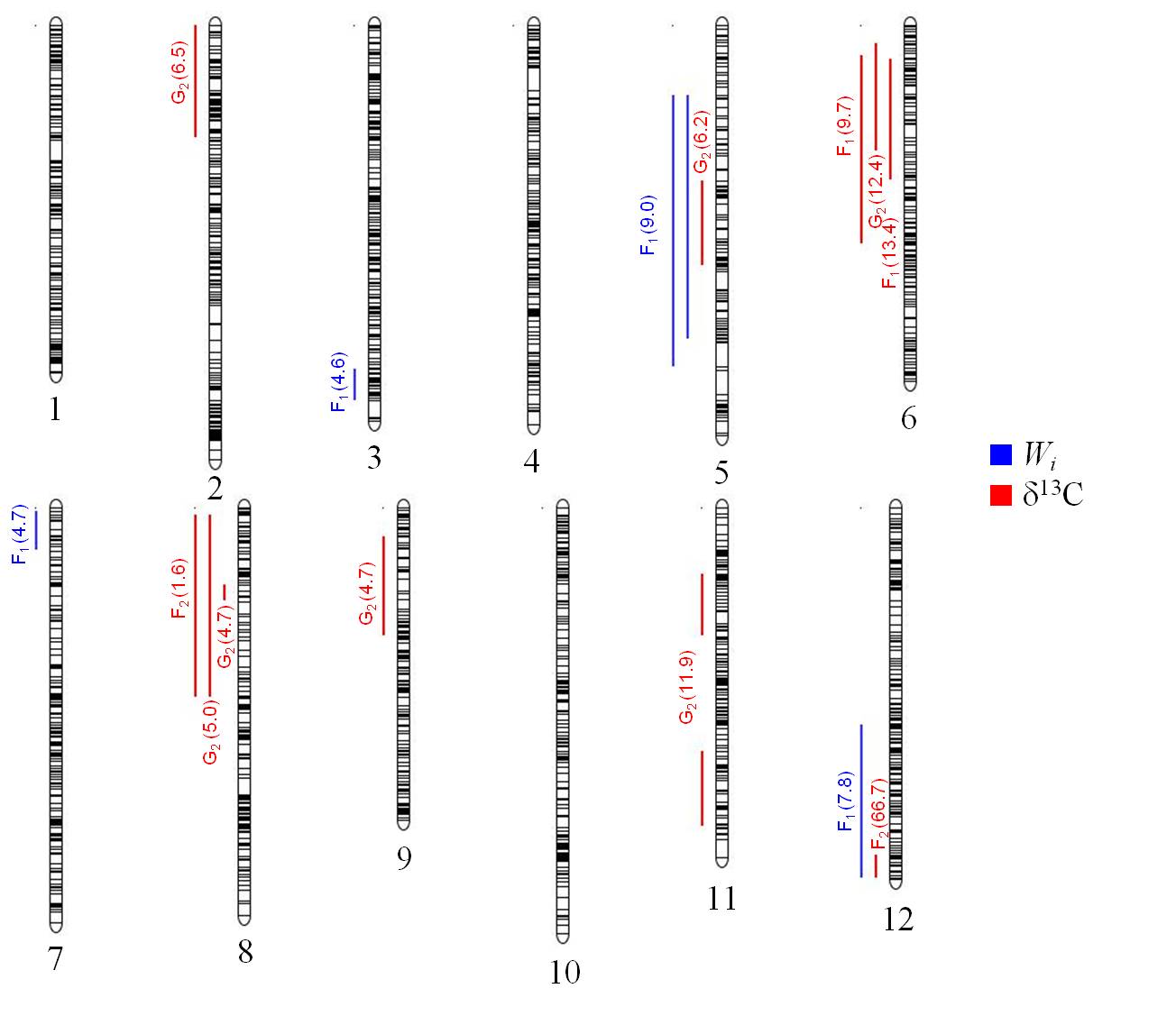

Figure 4. Map location of QTLs for intrinsic water-use efficiency (Wi) and carbon isotope composition (δ13C) on the composite linkage map of P. pinaster available from de Miguel et al. (2015). The QTLs shown were detected in three different pedigrees from independent studies: G2 (Brendel et al. 2002), F2 (Marguerit et al. 2014) and F1 (de Miguel et al. 2014). Percentage of explained phenotypic variance for each QTL is shown between brackets and 95% confidence intervals by vertical lines. QTLs were projected on the composite map with Biomercator v4.2.
Interestingly, genetic and QTL studies provide no evidence for a trade-off between WUE and growth in maritime pine, as these two traits are not genetically linked in the tested genetic background (Brendel et al., 2002; Marguerit et al., 2014). This suggests that it may be possible for forest tree breeders to improve WUE and growth at the same time. Studies of the genetic architecture of CR are also progressing, as in total 1,000 offspring from three pedigrees have been studied to date. In one pedigree (F2) in which more than 460 offspring were characterized, five QTLs for P50 were identified each accounting for 3% to 10% of the phenotypic variation (unpublished results): none colocalized with QTL for δ13C.
Linking genotype and environment, and molecular signatures of natural selection: identification of adaptive alleles
The identification of gene and gene networks involved in adaptation to environment is a key topic in ecological and evolutionary research. Several methods have been developed for identifying signatures of natural selection at the population level (reviewed by Rellstab et al., 2015) and the number of experimental studies on this topic, including some on non-model species such as forest trees, is increasing rapidly. One general difficulty associated with these studies concerns the association of molecular signatures of adaptation to the environment with specific traits over and above general insight gleaned from gene annotation. The situation is even more difficult for conifers, in which the functions of many genes remain unknown. No specific candidate gene studies have yet been carried out for WUE and CR in maritime pine, but several candidate drought-response genes have been identified as potentially relevant to adaptation to climate.
Two general approaches have been used in maritime pine to identify loci that have been targeted by selection. These approaches are based on the analysis of allele frequencies either between populations or between individuals across space. They are based on simple principles. So-called ‘outlier tests’ are used to compare levels of genetic differentiation (generally FST) with a null distribution obtained by simulating explicit demographic scenarios or based on empirical information. The ‘environmental associations’ approach is based on the likely occurrence of alleles conferring an advantage under particular environmental conditions at high frequencies in populations in which those conditions are prevalent. Only one outlier-based study has been carried out for drought-responsive genes in maritime pine. Eveno et al. (2008) combined different outlier detection methods and identified two genes displaying higher levels of differentiation than expected between 10 populations from contrasting climates (PR-AGP4, encoding a putative arabinogalactan/proline-rich protein, and erd3, a gene encoding a putative methyltransferase with a SAM-binding domain), consistent with diversifying selection on these genes. Some populations displayed greater divergence (e.g. Tamrabta in Morocco) than others, potentially creating opportunities for restricted gene flow and local adaptation. Grivet et al. (2011) carried out an environmental association study using a data set similar to that used by Eveno et al. (2008), in which they assessed the correlation of candidate gene allele frequency with climatic variables while correcting for population genetic structure. Evidence of association with climatic variables was obtained for two genes, DHN5 (dehydrin) and 4CL (4-coumarate: CoA ligase). The 4CL gene was also found to be associated with climate in Aleppo pine, a sister species of maritime pine. Significant environmental associations pointed to a role for temperature, particularly if extreme, rather than precipitation, as the key driver of climatic adaptation in this species. This result was confirmed in a recent study (Jaramillo-Correa et al., 2015b) in which 16 of the 21 robust associations of SNP allele frequency with climate (involving 18 SNPs) related to temperature variables, whereas only five related to precipitation. The most relevant significant SNP-climate associations were two non-synonymous polymorphisms from a heat-stress transcription factor and a protein kinase gene, respectively, and a synonymous site from 4CL (already detected in a much smaller sample by Grivet et al. (2011)). For more specific studies associating candidate genes for WUE and CR with environment, the first step is identifying the relevant underlying molecular variant for these traits. Such studies are currently underway, and both phenotypic and genotypic information is being collected for a range-wide clonal collection (CLONAPIN, consisting of 512 clonally propagated genotypes from 35 populations representing the six distinct gene pools of maritime pine, https://www6.bordeaux-aquitaine.inra.fr/biogeco/Recherche/Sites-experimentaux/CLONAPIN) and a large regional collection of Corsican genotypes (900 half-sib families from 30 Corsican populations, https://www6.bordeaux-aquitaine.inra.fr/biogeco/Recherche/Sites-experimentaux/Ressources-genetiques-Pin-maritime-de-Corse).
Identifying drought-responsive genes/proteins and those important for adaptation, through transcriptomic and proteomic approaches
Finding Understanding the implication of genes and/or gene products involved in the expression of complex traits, such as WUE or CR, requires a profound knowledge of genome structure and function, which is still lacking for conifer species (De La Torre et al., 2014). However, considerable efforts have been made in recent decades to identify the genes regulated during adaptation to water deficit in forest trees. The first studies were mostly based on “low-throughput” approaches, such as cDNA-AFLP, microarray and qPCR. These techniques are not suitable for analyzing the entire transcriptomic response during abiotic stress, but they nevertheless led to the identification of genes displaying enhanced or repressed expression during the response to water deficit (reviewed by Cossu et al., 2014). The first studies in maritime pine used cDNA-AFLP to identify 48 DNA fragments differentially expressed in needles and roots (Dubos et al., 2003; Dubos and Plomion, 2003). Suppression-subtraction hybridization libraries have been analyzed with microarrays, with the results confirmed by RT-PCR, and this approach has led to the identification of more than 200 candidate genes for involvement in the water deficit response (Perdiguero et al., 2012a; 2013). These gene expression studies have highlighted the role of different processes during the acclimation of maritime pine to a period of long-lasting water deficit: growth inhibition, effects on carbohydrate metabolism, decreases in transpiration and photosynthesis and increases in cell rescue and plant defense. The general nature of the variation in response to water deficit is conserved in aerial organs and roots (Dubos et al., 2003), but the expression of some genes, the degree of variation and the time-dependence of the response may be organ-specific (Dubos and Plomion, 2001; Perdiguero et al., 2012a; 2013). Several studies have focused on the expression of genes encoding dehydrins, which play a major role in plant responses to abiotic stresses. These studies have suggested that the proteins of this complex family may play diverse specialized roles in response to water deprivation (Velasco-Conde et al., 2012; Perdiguero et al., 2012b; 2014). Genes encoding components of cuticular wax content and overexpressed in response to water deficit were identified by Le Provost et al. (2013), highlighting the importance of the non-stomatal control of transpiration as an adaptation mechanism in maritime pine. Metabolites and transcriptome profiling analyses have also been performed during the formation of earlywood and latewood. Using cDNA-AFLP techniques, Le Provost et al. (2003) identified about 100 genes displaying qualitative or quantitative variation during wood formation during the course of the growing season. Paiva et al. (2008) reported that water deficit during latewood formation caused an increase in carbon allocation to lignin biosynthesis, leading to an increase in cell wall thickening, enabling the tree to protect its hydraulic conductance. This study also provided a list of expressional candidate genes regulated during the course of the growing season and potentially involved in adaptation to water deficit. For example, genes related to transcription, stress responses and ABA signaling were found to be upregulated in unfavorable environmental conditions.
Next-generation sequencing technologies, which can yield millions of short reads in a single run, are becoming the method of choice for whole-transcriptome analysis during the drought stress response in forest trees. They provide a digital gene expression analysis that is much more precise than that achieved with older methods, particularly for genes with low levels of expression. These sequencing approaches are not yet widely used in forest trees, but they have been successfully used in poplar (Cossu et al., 2014), eucalyptus (Villar et al., 2011) and silver fir (Behringer et al., 2015), to decipher the molecular mechanisms underlying adaptation to water deficit. One recent study used RNAseq methods to analyze the whole-transcriptome response to environmental conditions in two maritime pine ecotypes (Cañas et al., 2015b). Differential expression was observed between populations from contrasting environments grown under the same conditions. The findings of this study highlighted the important role of transcriptome regulation in the environmental adaptation of maritime pine.
Fewer studies of metabolism and protein levels have been carried out to investigate the drought stress response in maritime pine. Early studies focusing on the analysis of a few metabolites revealed the importance of some mechanisms of tolerance to water deficit, such as osmotic adjustment (Nguyen and Lamant, 1988) and the accumulation of proline (Corcuera et al., 2012). More integrated metabolome studies have been recently published (Cañas et al., 2015a; de Miguel et al., 2016). The combination of high-throughput metabolomics applied to different plant organs revealed that different metabolic pathways are activated in aerial and underground organs in response to drought (de Miguel et al., 2016). Cañas et al., (2015a) combined the study of a large number of metabolites and co-expression network analysis under natural conditions. They showed that the environment regulates gene expression to fine-tune the metabolome during development. Protein accumulation in response to water deprivation and its genetic determinism have been also studied in maritime pine (Costa et al., 1998; Costa and Plomion, 1999). Nevertheless, studies with state-of-the art proteomics technology (Abril et al., 2011; Van Oudenhove and Devreese, 2013) would be required to validate the results obtained in gene expression studies and obtain a full understanding of the molecular response to drought in maritime pine.
Direct relationships between variations of temporal or spatial gene expression on the one hand, and DNA polymorphism on the other, should facilitate the identification of causal mutations potentially involved in modifying gene expression (cis-effects), but the integration of functional genomic studies to establish links between genotype and phenotype or to identify adaptive alleles remains rare for forest trees (but see Prunier et al., 2015). With the spread of transcriptome profiling based on deep-sequencing technologies (RNAseq), it has become possible to investigate variability in gene expression in large populations. This approach opens up new possibilities for improving our understanding of the way in which expression variation contributes to water deficit adaptation in maritime pine. Epigenetics studies should also provide us with a more detailed understanding of the contribution of gene expression to adaptive function. These areas have yet to be explored in this species.
Conclusion
Water and temperature are among the most critical environmental factors influencing the growth and development of plants. It has been predicted that water shortage will increase considerably by the end of the 21st century, in many regions of the globe. The Mediterranean basin will be particularly affected, with a predicted 3–4°C increase in mean annual temperature, a 10-20% decrease in annual precipitation and more frequent and severe drought periods (IPCC, 2014). It has also been predicted that extreme events, such as severe water deficit, will become more frequent (Beniston et al., 2007). These changes will have an impact on the fate of forest tree populations (in terms of seedling recruitment and survival, biomass production, susceptibility to fires and pathogens) in this region. As the regions in which maritime pine grows (i.e. France, Spain, Italy and Portugal and northern Africa) are likely to undergo major changes, increasing the survival and maintenance of growth in conditions of low water availability is a matter of major concern in this species.
Adaptation to climate change over an uncertain timescale will require a stock of ready-to-use options to decrease the risk of losses in the future. A number of possible solutions exist for influencing stand structure and composition and minimizing the adverse impacts of climate change. The valorization of within-species genetic variability for key adaptive traits will probably prove to be particularly important. This review describes the current state of knowledge regarding this question for maritime pine. The results obtained to date have made it possible to estimate basic genetic parameters for WUE and CR and to evaluate breeding opportunities. They have provided us with information about the evolutionary forces that have shaped phenotypic variability for these traits and have thus, improved our understanding of the maintenance and evolution of genetic variation in the wild. The advent of high-throughput genotyping technologies has also made it possible to determine how genotypic variation results in adaptive phenotypic variation and to develop new approaches for better predicting genetic performances and climate-driven shifts in the distribution of maritime pine.
New challenges are emerging. We need to reinforce the connections between i) functional genomics, which can be used to identify expressional candidate genes in response to water deficit, ii) population genomics, which identifies putative adaptive SNPs, iii) quantitative genetics, which provides clues to the way in which a phenotype is determined by its genotype, iv) evolutionary ecophysiology, which seeks to understand the micro-evolutionary forces responsible for shaping the phenotype, and v) ecological genomics, which considers the way in which the content of an organism’s genome affects its current distribution, possibly leading to the detection of SNPs associated with environmental variables. The relationships between these disciplines and the information they provide (e.g. adaptive DNA sequence polymorphisms, gene expression pattern, trait variability and their associations with the environment) have not yet been fully elucidated in maritime pine and the effort required to achieve such integration should not be underestimated (Prunier et al., 2016). Unlocking the adaptive potential of this species in the framework of climate change and associated biotic threats to ensure a sustainable production of wood is strategic. In this context, new paths to turn knowledge into the flow of breeding should be actively considered. This includes for the present case study, the development of proof of concepts to evaluate the integration of new genetic resources into the Aquitaine breeding population (e.g. in hybrid combinations), and the evaluation of the relevance of genomic prediction for key traits that are difficult or costly to measure (Bartholomé et al. 2016).
Acknowledgments: Over the last 10 years, research on these topics conducted at INRA-BIOGECO has received funding from INRA-EFPA division, ANR (SUSTAINPINE, FLAG and WUEtree projects), the European Commission (TREESNIP-QLK3-CT2002-01973, NOVELTREE-FP7-211868, PROCOGEN-FP7-289841), FEDER (ABIOGEN No. Presage 32973) and the ‘Investments for the Future’ program (ANR-10-EQPX-16, XYLOFOREST) of the French National Agency for Research. JB was supported by a postdoctoral fellowship from ‘Conseil Général des Landes’. HL held a PhD fellowship from INRA (EFPA division, ACCAF metaprogram) and Région Aquitaine (no. 20111203004). MdM held a postdoctoral fellowship from PROCOGEN and WUETree. EM was a visiting scientist at INRA-BIOGECO. We thank Monika Konnert and an anonymous reviewer for their careful reading of our manuscript and constructive comments.
Authors’ contributions: CP coordinated this review and defined its content. All authors contributed to the paper, read and approved the submitted manuscript.
References
- Abril N, Gion JM, Kerner R, Müller-Starck G, Cerrillo RMN, Plomion C, Renaut J, Valledor L, Jorrin-Novo JV. 2011. Proteomics research on forest trees, the most recalcitrant and orphan plant species. Phytochemistry 72: 1219-1242. doi:10.1016/j.phytochem.2011.01.005
- Alberto FJ, Aitken SN, Alía R, González-Martínez SC, Hänninen H, Kremer A, Lefèvre F, Lenormand T, Yeaman S, Whetten R, et al. 2013. Potential for evolutionary responses to climate change – evidence from tree populations. Global Change Biology 19: 1645-1661. doi:10.1111/gcb.12181
- Allen CD, Breshears DD, McDowell NG. 2015. On underestimation of global vulnerability to tree mortality and forest die-off from hotter drought in the Anthropocene. Ecosphere 6: art129. doi:10.1890/ES15-00203.1
- Allen CD, Macalady AK, Chenchouni H, Bachelet D, McDowell N, Vennetier M, Kitzberger T, Rigling A, Breshears DD, Hogg EH, et al. 2010. A global overview of drought and heat-induced tree mortality reveals emerging climate change risks for forests. Forest Ecology and Management 259: 660-684. doi:10.1016/j.foreco.2009.09.001
- Anderegg WR, Flint A, Huang CY, Flint L, Berry JA, Davis FW, Sperry JS, Field CB. 2015. Tree mortality predicted from drought-induced vascular damage. Nature Geoscience 8: 367–371. doi:10.1038/ngeo2400
- Aranda I, Alia R, Ortega U, Dantas, AK, Majada J. 2010. Intra-specific variability in biomass partitioning and carbon isotopic discrimination under moderate drought stress in seedlings from four Pinus pinaster populations. Tree Genetics and Genomes 6: 169-178. doi:10.1007/s11295-009-0238-5
- Bartholomé J, Van Heerwaarden J, Isik F, Boury C, Vidal M, Plomion C, Bouffier L. 2016. Performance of genomic prediction within and across generations in maritime pine. BMC Genomics(in press). doi:10.1186/s12864-016-2879-8
- Bartholomé J, Mandrou E, Mabiala A, Jenkins J, Nabihoudine I, Klopp C, J Schmutz, Plomion C, Gion JM. 2015a. High-resolution genetic linkage maps of Eucalyptus improve BRASUZ1 reference genome assembly. New Phytologist 206:1283-96. doi:10.1111/nph.13150
- Bartholomé J, Mabiala A, Savelli B, Bert D, Brendel O, Plomion C, Gion JM. 2015b. Genetic architecture of carbon isotope composition and growth in Eucalyptus across multiple environments. New Phytologist 206: 1437-1449. doi:10.1111/nph.13301
- Barton NH. 1999. Clines in polygenic traits. Genetics Research 74: 223-236. doi:10.1017/S001667239900422X
- Behringer D, Zimmermann H, Ziegenhagen B, Liepelt S. 2015. Differential gene expression reveals candidate genes for drought stress response in Abies alba (Pinaceae). PLoS ONE 10: e0124564. doi:10.1371/journal.pone.0124564
- Beniston M, Stephenson DB, Christensen OB, Ferro CAT, Frei C, Goyette S, Halsnaes K, Holt T, Jylhä K, Koffi B, et al. 2007. Future extreme events in European climate: an exploration of regional climate model projections. Climatic Change 81: 71-95. doi:10.1007/s10584-006-9226-z
- Bodénès C, Chancerel E, Ehremann F, Kremer A, Plomion C. 2016. High-density linkage mapping and distribution of segregation distortion regions in the oak genome. DNA Research (in press). doi:10.1093/dnares/dsw001
- Bogeat-Triboulot MB, Brosché M, Renaut J, et al. 2007. Gradual soil water depletion results in reversible changes of gene expression, protein profiles, ecophysiology, and growth performance in Populus euphratica, a poplar growing in arid regions. Plant Physiology 143: 876–92. doi:10.1104/pp.106.088708
- Bouche PS, Delzon S, Choat B, Badel E, Brodribb TJ, Burlett R, Cochard H, Charra-Vaskou K, Lavigne B, Li S, Mayr S, Morris H, Torres-Ruiz JM, Zufferey V, Jansen S. 2015. Are needles of Pinus pinaster more vulnerable to xylem embolism than branches? New insights from X-ray computed tomography. Plant Cell and Environment 39: 860–870. doi:10.1111/pce.12680
- Bouche PS, Jansen S, Sabalera JC, Cochard H, Burlett R, Delzon S. 2016. Low intra-tree variability in resistance to embolism in four Pinaceae species. Annals of Forest Science 73: 681-689. doi:10.1007/s13595-016-0553-6
- Bouche PF, Larter M, Domec JC, Burlett R, Gasson P, Jansen S, Delzon S. 2014. A broad survey of xylem hydraulic safety and efficiency in conifers. Journal of Experimental Botany 65: 4419-4431. doi:10.1093/jxb/eru218
- Breda N, Huc R, Granier A, Dreyer E. 2006. Temperate forest trees and stands under severe drought: a review of ecophysiological responses, adaptation processes and long-term consequences. Annals of Forest Science 63: 625-644. doi:10.1051/forest:2006042
- Brendel O, Pot D, Plomion C, Rozenberg P, Guehl JM. 2002. Genetic parameters and QTL analysis of δ13C and ring width in maritime pine. Plant, Cell and Environment 25: 945-953. doi:10.1046/j.1365-3040.2002.00872.x
- Brodribb TJ, Cochard H. 2009. Hydraulic failure defines the recovery and point of death in water-stressed conifers. Plant Physiology 149: 575-584. doi:10.1104/pp.108.129783
- Brodribb TJ, Feild TS, Sack L. 2010. Viewing leaf structure and evolution from a hydraulic perspective. Functional Plant Biology 37: 488-498. doi:10.1071/FP10010
- Budde KB, Heuertz M, Hernández-Serrano A, Pausas JG, Vendramin GG, Verdú M, González-Martínez SC. 2014. In situ genetic association for serotiny, a fire-related trait, in Mediterranean maritime pine (Pinus pinaster). New Phytologist 201: 230-241. doi:10.1111/nph.12483
- Canales J, Bautista R, Label P, Gómez-Maldonado J, Lesur I, Fernández-Pozo N, Rueda-López M, Guerrero-Fernández D, Castro-Rodríguez V, Benzekri H, et al. 2014. De novo assembly of maritime pine transcriptome: implications for forest breeding and biotechnology. Plant Biotechnology Journal 12: 286-299. doi:10.1111/pbi.12136
- Cañas RA, Canales J, Muñoz-Hernández C, Granados JM, Ávila C, García-Martín ML, Cánovas FM. 2015a. Understanding developmental and adaptive cues in pine through metabolite profiling and co-expression network analysis. Journal of Experimental Botany 66: 3113-3127. doi:10.1093/jxb/erv118
- Cañas RA, Feito I, Fuente-Maqueda J, Ávila C, Majada J, Cánovas F. 2015b. Transcriptome-wide analysis supports environmental adaptations of two Pinus pinaster populations from contrasting habitats. BMC Genomics 16: 1-17.
- Carle J, Holmgren P. 2008. Wood from planted forests - a global outlook 2005-2030. Forest Products Journal 58: 7.
- Chagné D, Brown G, Lalanne C, Madur D, Pot D, Neale D, Plomion C. 2003. Comparative genome and QTL mapping between maritime and loblolly pines. Molecular Breeding 12: 185-195. doi:10.1023/A:1026318327911
- Chancerel E, Lamy JB, Lesur I, Noirot C, Klopp C, Ehrenmann F, Boury C, Provost GL, Label P, Lalanne C, et al. 2013. High-density linkage mapping in a pine tree reveals a genomic region associated with inbreeding depression and provides clues to the extent and distribution of meiotic recombination. BMC Biology 11: 50. doi:10.1186/1741-7007-11-50
- Chancerel E, Lepoittevin C, Le Provost G, Lin YC, Jaramillo-Correa J, Eckert A, Wegrzyn J, Zelenika D, Boland A, Frigerio JM, et al. 2011. Development and implementation of a highly-multiplexed SNP array for genetic mapping in maritime pine and comparative mapping with loblolly pine. BMC Genomics 12: 368. doi:10.1186/1471-2164-12-368
- Choat B, Jansen S, Brodribb TJ, Cochard H, Delzon S, Bhaskar R, Bucci SJ, Feild TS, Gleason SM, Hacke UG, et al. 2012. Global convergence in the vulnerability of forests to drought. Nature 491: 752-755. doi:10.1038/nature11688
- Ciais P, Reichstein M, Viovy N, Granier A, Ogee J, Allard V, Aubinet M, Buchmann N, Bernhofer C, Carrara A, et al. 2005. Europe-wide reduction in primary productivity caused by the heat and drought in 2003. Nature 437: 529-533. doi:10.1038/nature03972
- Cochard H, Badel E, Herbette S, Delzon S, Choat B, Jansen S. 2013. Methods for measuring plant vulnerability to cavitation: a critical review. Journal of Experimental Botany 64: 4779–4791. doi:10.1093/jxb/ert193
- Cochard H, Damour G, Bodet C, Tharwat I, Poirier M, Améglio T. 2005. Evaluation of a new centrifuge technique for rapid generation of xylem vulnerability curves. Physiologia Plantarum 124: 410-41. doi:10.1111/j.1399-3054.2005.00526.x
- Cochard H, Barigah ST, Kleinhentz M , Eshel A. 2008. Is xylem cavitation resistance a relevant criterion for screening drought resistance amongst Prunus species ? Journal of Plant Physiology 165: 976-982. doi:10.1016/j.jplph.2007.07.020
- Coop G, Witonsky D, Di Rienzo A, Pritchard JK. 2010. Using environmental correlations to identify loci underlying local adaptation. Genetics 185: 1411-1423. doi:10.1534/genetics.110.114819
- Corcuera L, Cochard H, Gil-Pelegrin E, Notivol E. 2011. Phenotypic plasticity in mesic populations of Pinus pinaster improves resistance to xylem embolism (P50) under severe drought. Trees 25: 1033-1042. doi:10.1007/s00468-011-0578-2
- Corcuera L, Gil-Pelegrin E, Notivol E. 2012. Aridity promotes differences in proline and phytohormone levels in Pinus pinaster populations from contrasting environments. Trees 26:799–808. doi:10.1007/s00468-011-0651-x
- Corcuera L, Gil-Pelegrin E, Notivol E. 2010. Phenotypic plasticity in Pinus pinaster δ13C: environment modulates genetic variation. Annals of Forest Science 67: 812–812. doi:10.1051/forest/2010048
- Cossu R, Giordani T, Cavallini A, Natali L. 2014. High-throughput analysis of transcriptome variation during water deficit in a poplar hybrid: a general overview. Tree Genetics and Genomes 10: 53-66. doi:10.1007/s11295-013-0661-5
- Costa P, Bahrman N, Frigerio J-M, Kremer A, Plomion C. 1998. Water-deficit-responsive proteins in maritime pine. Plant Molecular Biology 38: 587-596. doi:10.1023/A:1006006132120
- Costa P, Plomion C. 1999. Genetic analysis of needle proteins in maritime pine 2. Variation of protein accumulation. Silvae Genetica 48: 3-4.
- De La Torre AR, Birol I, Bousquet J, Ingvarsson PK, Jansson S, Jones SJN, Keeling CI, MacKay J, Nilsson O, Ritland K, et al. 2014. Insights into conifer giga-genomes. Plant Physiology 166: 1724-1732. doi:10.1104/pp.114.248708
- de Miguel M, Guevara MA, Sánchez-Gómez D, de María N, Díaz LM, Mancha JA, Fernández de Simón B, Cadahía E, Desai N, Aranda I, Cervera MT. 2016. Organ-specific metabolic responses to drought in Pinus pinaster Ait. Plant Physiology and Biochemistry 102: 17-26. doi:10.1016/j.plaphy.2016.02.013
- de Miguel M, Bartholomé J, Ehrenmann F, Murat F, Moriguchi Y, Uchiyama K, Ueno S, Tsumura Y, Lagraulet H, de Maria N, et al. 2015. Evidence of intense chromosomal shuffling during conifer evolution. Genome Biology and Evolution 7: 2799-2809. doi:10.1093/gbe/evv185
- de Miguel M, Cabezas JA, de Maria N, Sanchez-Gomez D, Guevara MA, Velez MD, Saez-Laguna E, Diaz LM, Mancha JA, Barbero MC, et al. 2014. Genetic control of functional traits related to photosynthesis and water use efficiency in Pinus pinaster Ait. drought response: integration of genome annotation, allele association and QTL detection for candidate gene identification. BMC Genomics 15: 464. doi:10.1186/1471-2164-15-464
- de Miguel M, Sánchez-Gómez D, Cervera MT, Aranda I. 2012. Functional and genetic characterization of gas exchange and intrinsic water use efficiency in a full-sib family of Pinus pinaster Ait. in response to drought. Tree Physiology 32: 94-103. doi:10.1093/treephys/tpr122
- De Swaef T, De Schepper V, Vandegehuchte MW, Steppe K. 2015. Stem diameter variations as a versatile research tool in ecophysiology. Tree Physiology 35:1047-1061. doi:10.1093/treephys/tpv080
- Delzon S, Cochard H. 2014. Recent advances in tree hydraulics highlight the ecological significance of the hydraulic safety margin. New Phytologist 203: 355-358. doi:10.1111/nph.12798
- Delzon S, Douthe C, Sala A, Cochard H. 2010. Mechanism of water-stress induced cavitation in conifers: bordered pit structure and function support the hypothesis of seal capillary-seeding. Plant Cell and Environment 33: 2101–2111. doi:10.1111/j.1365-3040.2010.02208.x
- Denis JB. 1988. Two way analysis using covariates. Statistics 19: 123-132. doi:10.1080/02331888808802080
- Deschamps S, Llaca V, May GD. 2012. Genotyping-by-sequencing in plants. Biology 1: 460. doi:10.3390/biology1030460
- Drew D, Downes G, Evans R. 2011. Short-term growth responses and associated wood density fluctuations in variously irrigated Eucalyptus globulus. Trees - Structure and Function 25: 153-161.
- Drew DM, Downes GM. 2009. The use of precision dendrometers in research on daily stem size and wood property variation: A review. Dendrochronologia 27: 159-172. doi:10.1016/j.dendro.2009.06.008
- Dubos C, Le Provost G, Pot D, Salin F, Lalane C, Madur D, Frigerio JM, Plomion C. 2003. Identification and characterization of water-stress-responsive genes in hydroponically grown maritime pine (Pinus pinaster) seedlings. Tree Physiology 23: 169-179. doi:10.1093/treephys/23.3.169
- Dubos C, Plomion C. 2001. Drought differentially affects expression of a PR-10 protein, in needles of maritime pine (Pinus pinaster Ait.) seedlings. Journal of Experimental Botany 52: 1143-1144. doi:10.1093/jexbot/52.358.1143
- Dubos C, Plomion C. 2003. Identification of water-deficit responsive genes in maritime pine (Pinus pinaster Ait.) roots. Plant Molecular Biology 51: 249-262. doi:10.1023/A:1021168811590
- Dvorak WS. 2012. Water use in Plantations of eucalypts and pines: a discussion paper from a tree breeding perspective. International Forestry Review 14: 110–119. doi:10.1505/146554812799973118
- Dzwonko Z. 2001. Assessment of light and soil conditions in ancient and recent woodlands by Ellenberg indicator values. Journal of Applied Ecology 38: 942-951. doi:10.1046/j.1365-2664.2001.00649.x
- Ekblom R, Galindo J. 2011. Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity 107: 1-15. doi:10.1038/hdy.2010.152
- Evans LM, Slavov GT, Rodgers-Melnick E, Martin J, Ranjan P, Muchero W, Brunner AM, Schackwitz W, Gunter L, Chen JG, et al. 2014. Population genomics of Populus trichocarpa identifies signatures of selection and adaptive trait associations. Nature Genetics 46: 1089-1096. doi:10.1038/ng.3075
- Eveno E, Collada C, Guevara MA, Léger V, Soto A, Díaz L, Léger P, González-Martínez SC, Cervera MT, Plomion C, et al. 2008. Contrasting patterns of selection at Pinus pinaster Ait. drought stress candidate genes as revealed by genetic differentiation analyses. Molecular Biology and Evolution 25: 417-437. doi:10.1093/molbev/msm272
- FAO 2012. FRA 2015 - Terms and definitions. Forest resources Assessment Working Paper 180. Rome: Food and Agriculture Organization of the United Nations. 31.
- Farquhar GD, Ehleringer JR, Hubick KT. 1989. Carbon isotope discrimination and photosynthesis. Annual Review of Plant Physiology and Plant Molecular Biology 40: 503–537. doi:10.1146/annurev.pp.40.060189.002443
- Fernández M, Gil L, Pardos JA. 1999. Response of Pinus pinaster Ait. provenances at early age to water supply. I. Water relation parameters. Annals of Forest Science 56: 179-187. doi:10.1051/forest:19990209
- Fernández M, Gil L, Pardos JA. 2000. Effects of water supply on gas exchange in Pinus pinaster Ait. provenances during their first growing season. Annals of Forest Science 57: 9-16. doi:10.1051/forest:2000107
- Fichot R, Barigah TS, Chamaillard S, Le Thiec D, Laurans F, Cochard H, Brignolas F. 2010. Common trade-offs between xylem resistance to cavitation and other physiological traits do not hold among unrelated Populus deltoides×Populus nigra hybrids. Plant Cell and Environment 33: 1553-1568. doi:10.1111/j.1365-3040.2010.02164.x
- Frichot E, Schoville SD, Bouchard G, François O. 2013. Testing for associations between loci and environmental gradients using latent factor mixed models. Molecular Biology and Evolution 30: 1687-1699. doi:10.1093/molbev/mst063
- Furbank RT, Tester M. 2011. Phenomics – technologies to relieve the phenotyping bottleneck. Trends in Plant Science 16: 635-644. doi:10.1016/j.tplants.2011.09.005
- Galmés J, Flexas J, Savé R, Medrano H (2007) Water relations and stomatal characteristics of Mediterranean plants with different growth forms and leaf habits: responses to water stress and recovery. Plant and Soil 290: 139–155. doi:10.1007/s11104-006-9148-6
- Gnirke A, Melnikov A, Maguire J, Rogov P, LeProust EM, Brockman W, Fennell T, Giannoukos G, Fisher S, Russ C, et al. 2009. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nature Biotechechnology 27: 182-189. doi:10.1038/nbt.1523
- Granier A, Loustau D. 1994. Measuring and modelling the transpiration of a maritime pine canopy from sap-flow data. Agricultural and Forest Meteorology 71: 61-81. doi:10.1016/0168-1923(94)90100-7
- Grivet D, Climent J, Zabal-Aguirre M, Neale DB, Vendramin GG, González-Martínez SC. 2013. Adaptive evolution of Mediterranean pines. Molecular Phylogenetics and Evolution 68: 555-566. doi:10.1016/j.ympev.2013.03.032
- Grivet D, Sebastiani F, Alía R, Bataillon T, Torre S, Zabal-Aguirre M, Vendramin GG, González-Martínez SC. 2011. Molecular footprints of local adaptation in two Mediterranean conifers. Molecular Biology and Evolution 28: 101-116. doi:10.1093/molbev/msq190
- Guehl J-M, Nguyen-Queyrens A, Loustau D, Ferhi A. 1995 Genetic and environmental determinants of water-use efficiency and carbon isotope discrimination in forest trees. In: Sandermann H, Bonnet-Masimbert M (eds) Eurosilva Contrib. to For. tree Physiol. Results from Eurosilva Proj. Present. Dourdan, Fr. 7-10 Novemb. 1994. Editions Colloques de l’INRA, Paris, pp 297–321.
- Hacke UG, Sperry JS, Pockman WT, Davis SD, McCulloh KA. 2001. Trends in wood density and structure are linked to prevention of xylem implosion by negative pressure. Oecologia, 126: 457-461. doi:10.1007/s004420100628
- Hartmann H. 2015. Carbon starvation during drought-induced tree mortality – are we chasing a myth? Journal of Plant Hydraulics 2: e-005. doi:10.20870/jph.2015.e005
- Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A. 2005. Very high resolution interpolated climate surfaces for global land areas. International Journal of Climatology 25: 1965-1978. doi:10.1002/joc.1276
- Hunault G, Gaillard S, Bourbeillon J, Montiel G, Simier P, Campion C, Jacques M-A, Belin E, Boureau T. 2015. Phenoplant: a web resource for the exploration of large chlorophyll fluorescence image datasets. Plant Methods 11: 1-12.
- Illy G. 1966. Recherches sur l'amélioration génétique du pin maritime. Annales des Sciences Forestières. 23: 765-948. doi:10.1051/forest/19660401
- IPCC Climate Change 2014: Impacts, Adaptation, and Vulnerability. Part A: Global and Sectoral Aspects. Contribution of Working Group II to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change [Field CB, Barros VR, Dokken DJ, Mach KJ, Mastrandrea MD, Bilir TE, Chatterjee M, Ebi KL, Estrada YO, Genova RC, Girma B, Kissel ES, Levy AN, MacCracken S, Mastrandrea PR, White LL (eds.)]. Cambridge University Press, Cambridge, United Kingdom and New York, NY, USA, 1132 pp.
- Isik F, Bartholomé J, Farjat A, Chancerel E, Raffin A, Sanchez L, Plomion C, Bouffier L. 2015. Genomic selection in maritime pine. Plant Science 242: 108-119. doi:10.1016/j.plantsci.2015.08.006
- Jaramillo-Correa JP, Prunier J, Vázquez-Lobo A, Keller SR, Moreno-Letelier A. 2015a. Molecular signatures of adaptation and selection in forest trees. In: Pomion C, Adam-Blondon AF eds. Advances in Botanical Research: Academic Press, 265-306. doi:10.1016/bs.abr.2015.04.003
- Jaramillo-Correa JP, Rodríguez-Quilón I, Grivet D, Lepoittevin C, Sebastiani F, Heuertz M, Garnier-Géré PH, Alía R, Plomion C, Vendramin GG, et al. 2015b. Molecular proxies for climate maladaptation in a long-lived tree (Pinus pinaster Aiton, Pinaceae). Genetics 199: 793-807. doi:10.1534/genetics.114.173252
- King G, Fonti P, Nievergelt D, Büntgen U, Frank D. 2013. Climatic drivers of hourly to yearly tree radius variations along a 6°C natural warming gradient. Agricultural and Forest Meteorology 168: 36-46. doi:10.1016/j.agrformet.2012.08.002
- Lagraulet H. 2015. Plasticité phénotypique et architecture génétique de la croissance et de la densité du bois du pin maritime (Pinus pinaster Ait.). Université de Bordeaux France.
- Lamy JB, Bouffier L, Burlett R, Plomion C, Cochard H, Delzon S. 2011. Uniform selection as a primary force reducing population genetic differentiation of cavitation resistance across a species range. PLoS ONE 6: e23476. doi:10.1371/journal.pone.0023476
- Lamy JB, Delzon S, Bouche PS, Alia R, Vendramin GG, Cochard H, Plomion C. 2014. Limited genetic variability and phenotypic plasticity detected for cavitation resistance in a Mediterranean pine. New Phytologist 201: 874-886. doi:10.1111/nph.12556
- Le Provost G, Paiva J, Pot D, Brach J, Plomion C. 2003. Seasonal variation in transcript accumulation in wood forming tissues of maritime pine (Pinus pinaster Ait) with emphasis on a cell wall Glycine Rich Protein. Planta 217: 820-830. doi:10.1007/s00425-003-1051-2
- Le Provost G, Domergue F, Lalanne C, Ramos Campos P, Grosbois A, Bert D, Meredieu C, Danjon F, Plomion C, Gion JM. 2013. Soil water stress affects both cuticular wax content and cuticle-related gene expression in young saplings of maritime pine (Pinus pinaster Ait). BMC Plant Biology 13: 1-12. doi:10.1186/1471-2229-13-95
- Lepoittevin C, Frigerio JM, Garnier-Géré P, Salin F, Cervera M-T, Vornam B, Harvengt L, Plomion C. 2010. In vitro v.s in silico detected SNPs for the development of a genotyping array: what can we learn from a non-model species? PLoS ONE 5: e11034. doi:10.1371/journal.pone.0011034
- Lepoittevin C, Harvengt L, Plomion C, Garnier-Géré P. 2012. Association mapping for growth, straightness and wood chemistry traits in the Pinus pinaster Aquitaine breeding population. Tree Genetics and Genomes 8: 113-126. doi:10.1007/s11295-011-0426-y
- Lindner M, Maroschek M, Netherer S, Kremer A, Barbati A, Garcia-Gonzalo J, Seidl R, Delzon S, Corona P, Kolström M, et al. 2010. Climate change impacts, adaptive capacity, and vulnerability of European forest ecosystems. Forest Ecology and Management 259: 698-709. doi:10.1016/j.foreco.2009.09.023
- Loustau D, Bosc A, Colin A, Ogée J, Davi H, François C, Dufrêne E, Déqué M, Cloppet E, Arrouays D, et al. 2005. Modeling climate change effects on the potential production of French plains forests at the sub-regional level. Tree Physiology 25: 813-823. doi:10.1093/treephys/25.7.813
- MacDicken KG. 2015. Global Forest Resources Assessment 2015: What, why and how? Forest Ecology and Management 352: 3-8. doi:10.1016/j.foreco.2015.02.006
- Mackay J, Dean JD, Plomion C, Peterson D, Cánovas F, Pavy N, Ingvarsson P, Savolainen O, Guevara MÁ, Fluch S, et al. 2012. Towards decoding the conifer giga-genome. Plant Molecular Biology 80: 555-569. doi:10.1007/s11103-012-9961-7
- Maherali, H, Pockman WT, Jackson RB. 2004. Adaptive variation in the vulnerability of woody plants to xylem cavitation. Ecology 85: 2184-2199. doi:10.1890/02-0538
- Marguerit E, Bouffier L, Chancerel E, Costa P, Lagane F, Guehl JM, Plomion C, Brendel O. 2014. The genetics of water-use efficiency and its relation to growth in maritime pine. Journal of Experimental Botany 65: 4757-4768. doi:10.1093/jxb/eru226
- Mauricio R. 2001. Mapping quantitative trait loci in plants: uses and caveats for evolutionary biology. Nature Reviews Genetics 2: 370-381. doi:10.1038/35072085
- Meinzer FC, Ingamells JL, Crisosto C. 1991. Carbon isotope discrimination correlates with bean yield of diverse coffee popultions. HortScience 26: 1413–1414.
- Michelozzi M, Loreto F, Colom R, Rossi F, Calamassi R. 2011. Drought responses in Aleppo pine seedlings from two wild provenances with different climatic features. Photosynthetica 49: 564-572. doi:10.1007/s11099-011-0068-1
- Monclus R, Leple JC, Bastien C, Bert PF, Villar M, Marron N, Brignolas F, Jorge V. 2012. Integrating genome annotation and QTL position to identify candidate genes for productivity, architecture and water-use efficiency in Populus spp. BMC Plant Biology 12: 173. doi:10.1186/1471-2229-12-173
- Morgenstern M. 2011. Geographic variation in forest trees: genetic basis and application of knowledge in silviculture. Vancouver, BC: UBC Press.
- Munns R, James RA, Sirault XRR, Furbank RT, Jones HG. 2010. New phenotyping methods for screening wheat and barley for beneficial responses to water deficit. Journal of Experimental Botany 61: 3499-3507. doi:10.1093/jxb/erq199
- Neale DB, Savolainen O. 2004. Association genetics of complex traits in conifers. Trends in Plant Science 9: 325-330. doi:10.1016/j.tplants.2004.05.006
- Nguyen A, Lamant A. 1988. Pinitol and myo-inositol accumulation in water-stressed seedlings of maritime pine. Phytochemistry 27: 3423-3427. doi:10.1016/0031-9422(88)80742-8
- Nguyen-Queyrens A, Ferhi A, Loustau D, Guehl JM. 1998. Within-ring d13C spatial variability and interannual variation in wood cellulose of two contrasting provenances of Pinus pinaster. Canadian Journal Forest Research 28: 766-773.
- Nguyen A, Lamant A. 1989. Variation in growth and osmotic regulation of roots of water-stressed maritime pine (Pinus pinaster Ait.) provenances. Tree Physiology 5: 123-133. doi:10.1093/treephys/5.1.123
- Nguyen-Queyrens A, Bouchet-Lannat F. 2003. Osmotic adjustment in three-year-old seedlings of five provenances of maritime pine (Pinus pinaster) in response to drought. Tree Physiology 23: 397-404. doi:10.1093/treephys/23.6.397
- Nystedt B, Street NR, Wetterbom A, Zuccolo A, Lin YC, Scofield DG, Vezzi F, Delhomme N, Giacomello S, Alexeyenko A, et al. 2013. The Norway spruce genome sequence and conifer genome evolution. Nature 497: 579-584. doi:10.1038/nature12211
- Paiva J, Garnier-Géré PH, Rodrigues JC, Alves A, Santos S, Graça J, Le Provost G, Chaumeil P, da Silva-Perez D, Bosc A, Fevereiro P, Plomion C. 2008. Plasticity of maritime pine (Pinus pinaster Ait) wood forming tissues during a growing season. New Phytol 179: 1080-1094. doi:10.1111/j.1469-8137.2008.02536.x
- Papadakis JS. 1984. Advances in the analysis of field experiments. Proceedings of the Academy of Athens 59: 326–342.
- Pavy N, Pelgas B, Laroche J, Rigault P, Isabel N, Bousquet J. 2012. A spruce gene map infers ancient plant genome reshuffling and subsequent slow evolution in the gymnosperm lineage leading to extant conifers. BMC Biology 10: 1-19. doi:10.1186/1741-7007-10-84
- Payn T, Carnus JM, Freer-Smith P, Kimberley M, Kollert W, Liu S, Orazio C, Rodriguez L, Silva LN, Wingfield MJ. 2015. Changes in planted forests and future global implications. Forest Ecology and Management 352: 57-67. doi:10.1016/j.foreco.2015.06.021
- Pedro-Monzonís M, Solera A, Ferrer J, Estrela T, Paredes-Arquiola J. 2015. A review of water scarcity and drought indexes in water resources planning and management. Journal of Hydrology 527: 482-93. doi:10.1016/j.jhydrol.2015.05.003
- Perdiguero P, Collada C, Soto Á. 2014. Novel dehydrins lacking complete K-segments in Pinaceae. The exception rather than the rule. Frontiers in Plant Science 5: 682. doi:10.3389/fpls.2014.00682
- Perdiguero P, Barbero MdC, Cervera MT, Collada C, Soto Á. 2013. Molecular response to water stress in two contrasting Mediterranean pines (Pinus pinaster and Pinus pinea). Plant Physiology and Biochemistry 67: 199-208. doi:10.1016/j.plaphy.2013.03.008
- Perdiguero P, Collada C, Barbero MdC, Casado GG, Cervera MT, Soto A. 2012a. Identification of water stress genes in Pinus pinaster Ait. by controlled progressive stress and suppression-subtractive hybridization. Plant Physiology and Biochemistry 50:44–53. doi:10.1016/j.plaphy.2011.09.022
- Perdiguero P, Barbero MC, Cervera MT, Soto Á, Collada C. 2012b. Novel conserved segments are associated with differential expression patterns for Pinaceae dehydrins. Planta 236: 1863–1874. doi:10.1007/s00425-012-1737-4
- Picon C, Guehl JM, Ferhi A. 1996. Leaf gas exchange and carbon isotope composition responses to drought in a drought-avoiding (Pinus pinaster) and a drought-tolerant (Quercus petraea) species under present and elevated atmospheric CO2 concentrations. Plant Cell and Environment 19: 182-190. doi:10.1111/j.1365-3040.1996.tb00239.x
- Pittermann, J, Choat B, Jansen S, Stuart SA, Lynn L, Dawson TE. 2010. The relationships between xylem safety and hydraulic efficiency in the Cupressaceae: the evolution of pit membrane form and function. Plant Physiology 153, 1919-1931. doi:10.1104/pp.110.158824
- Plomion C, Bartholomé J, Lesur I, Boury C, Rodríguez-Quilón I, Lagraulet H, Ehrenmann F, Bouffier L, Gion JM, Grivet D, et al. 2016. High-density SNP assay development for genetic analysis in maritime pine (Pinus pinaster). Molecular Ecology Resources 16: 574-587. doi:10.1111/1755-0998.12464
- Plomion C, Durel CE, O'Malley DM. 1996a. Genetic dissection of height in maritime pine seedlings raised under accelerated growth conditions. Theoretical and Applied Genetics 93: 849-858. doi:10.1007/BF00224085
- Plomion C, Yani A, Marpeau A. 1996b. Genetic determinism of delta3-carene in maritime pine using RAPD markers. Genome 39: 1123-1127. doi:10.1139/g96-141
- Pot D, McMillan L, Echt C, Le Provost G, Garnier-Géré P, Cato S, Plomion C. 2005. Nucleotide variation in genes involved in wood formation in two pine species. New Phytologist 167: 101-112. doi:10.1111/j.1469-8137.2005.01417.x
- Pot D, Rodrigues J-C, Rozenberg P, Chantre G, Tibbits J, Cahalan C, Pichavant F, Plomion C. 2006. QTLs and candidate genes for wood properties in maritime pine (Pinus pinaster Ait.). Tree Genetics and Genomes 2: 10-24. doi:10.1007/s11295-005-0026-9
- Prunier J, Verta JP, MacKay JJ. 2016. Conifer genomics and adaptation: at the crossroads of genetic diversity and genome function. New Phytologist 209: 44-62. doi:10.1111/nph.13565
- Prunier J, Tessier G, Bousquet J, MacMay J. 2015. From genotypes to phenotypes: expression levels of genes encompassing adaptive SNPs in black spruce. Plant Cell Reports 34: 2111-2125. doi:10.1007/s00299-015-1855-0
- Ramírez-Valiente JA, Koehler K, Cavender-Bares J. 2015. Climatic origins predict variation in photoprotective leaf pigments in response to drought and low temperatures in live oaks (Quercus series Virentes). Tree Physiology 35: 521-534. doi:10.1093/treephys/tpv032
- Reinhardt K, Castanha C, Germino MJ, Kueppers LM. 2011. Ecophysiological variation in two provenances of Pinus flexilis seedlings across an elevation gradient from forest to alpine. Tree Physiology 31: 615-625. doi:10.1093/treephys/tpr055
- Rellstab C, Gugerli F, Eckert AJ, Hancock AM, Holderegger R. 2015. A practical guide to environmental association analysis in landscape genomics. Molecular Ecology 24: 4348-4370. doi:10.1111/mec.13322
- Richardson DM, Hellmann JJ, McLachlan JS, Sax DF, Schwartz MW, Gonzalez P, Jean Brennan E, Camacho A, Root TL, Sala OL et al. 2009. Multidimensional evaluation of managed relocation. Proceedings of the National Academy of Sciences of the United States of America 106: 9721–9724. doi:10.1073/pnas.0902327106
- Savolainen O, Pyhäjärvi T, Knürr T. 2007. Gene flow and local adaptation in trees. Annual Review of Ecology Evolution and Systematics 38: 595-619. doi:10.1146/annurev.ecolsys.38.091206.095646
- Settele J, Scholes R, Betts R, Bunn SE, Leadley P, Nepstad D, Overpeck JT, Taboada MA. 2014. Terrestrial and inland water systems. In: Field CB, Barros VR, Dokken DJ, Mach KJ, Mastrandrea MD, Bilir TE, Chatterjee M, Ebi KL, Estrada YO, Genova RC, Girma B, Kissel ES, Levy AN, MacCracken S, Mastrandrea PR, White LL eds. Climate Change 2014: Impacts, Adaptation, and Vulnerability. Part A: Global and Sectoral Aspects. Contribution of Working Group II to the Fifth Assessment Report of the Intergovernmental Panel of Climate Change. Cambridge, United Kingdom and New York, NY, USA: Cambridge University Press, 271-359.
- Steppe K, De Pauw DJW, Lemeur R. 2008. Validation of a dynamic stem diameter variation model and the resulting seasonal changes in calibrated parameter values. Ecological Modelling 218: 247-259. doi:10.1016/j.ecolmodel.2008.07.006
- Szymura TH, Szymura M, Macioł A. 2014. Bioindication with Ellenberg's indicator values: A comparison with measured parameters in Central European oak forests. Ecological Indicators 46: 495-503. doi:10.1016/j.ecolind.2014.07.013
- Tognetti R, Michelozzi M, Lauteri M, Brugnoli E Giannini R. 2000. Geographic variation in growth, carbon isotope discrimination, and monoterpene composition in Pinus pinaster Ait. provenances. Canadian Journal Forest Research 30: 1682-1690. doi:10.1139/x00-096
- Tringe SG, Rubin EM. 2005. Metagenomics: DNA sequencing of environmental samples. Nature Reviews Genetics 6: 805-814. doi:10.1038/nrg1709
- Tyree MT, Zimmermann MH. 2002. Hydraulic architecture of whole plants and plant performance. In Xylem Structure and the Ascent of Sap (pp. 175-214). Springer Berlin Heidelberg. doi:10.1007/978-3-662-04931-0_6
- Van Oudenhove L, Devreese B. 2013. A review on recent developments in mass spectrometry instrumentation and quantitative tools advancing bacterial proteomics. Applied Microbiology and Biotechnology 97: 4749-4762. doi:10.1007/s00253-013-4897-7
- Velasco-Conde T, Yakovlev I, Majada J, Aranda I, Johnsen Ø. 2012. Dehydrins in maritime pine (Pinus pinaster) and their expression related to drought stress response. Tree Genetics and Genomes 8: 957-973. doi:10.1007/s11295-012-0476-9
- Vicente-Serrano SM, Beguería S, Lorenzo-Lacruz J, Camarero JJ, López-Moreno JI, Azorin-Molina C, Revuelto J, Morán-Tejeda E, Sanchez-Lorenzo A. 2012. Performance of drought indices for ecological, agricultural, and hydrological applications. Earth Interactions 16: 1-27. doi:10.1175/2012EI000434.1
- Vieira J, Campelo F, Rossi S, Carvalho A, Freitas H, Nabais C. 2015. Adjustment capacity of maritime pine cambial activity in drought-prone environments. PLoS ONE 10: e0126223. doi:10.1371/journal.pone.0126223
- Villar E, Klopp C, Noirot C, Novaes E, Kirst M, Plomion C, Gion JM. 2011. RNA-Seq reveals genotype-specific molecular responses to water deficit in eucalyptus. BMC Genomics 12: 1-18. doi:10.1186/1471-2164-12-538
- Visscher PM, Hill WG, Wray NR. 2008. Heritability in the genomics era - concepts and misconceptions. Nature Reviews Genetics 9: 255-266. doi:10.1038/nrg2322
- Wahid N, González-Martínez SC, Hadrami IE, Boulli A. 2006. Variation of morphological traits in natural populations of maritime pine (Pinus pinaster Ait.) in Morocco. Annals of Forest Science. 63: 83-92. doi:10.1051/forest:20050100
- Wamelink GWW, Joosten V, van Dobben HF, Berendse F. 2002. Validity of Ellenberg indicator values judged from physico-chemical field measurements. Journal of Vegetation Science 13: 269-278. doi:10.1111/j.1654-1103.2002.tb02047.x
- WWF, IIASA. 2012. Living Forests Report. Gland, Switzerland.
- Zweifel R, Eugster W, Etzold S, Dobbertin M, Buchmann N, Häsler R. 2010. Link between continuous stem radius changes and net ecosystem productivity of a subalpine Norway spruce forest in the Swiss Alps. New Phytologist 187: 819-830. doi:10.1111/j.1469-8137.2010.03301.x
Attachments
Other
List of the main experimental trials studying the genetics of adaptation to water deficit in maritime pine.
DownloadOther
Examples of SNP clusters obtained in the F2 mapping population used by Plomion et al. (2015) with GenomeStudio software (Illumina). X-axis: normalized Theta values ranged from 0 (homozygous for one allele) to 1 (homozygous for the alternative allele). Y-axis: normalized sum of fluorescence intensities for the two fluorochromes used in the Illumina Infinium assay. Panel A shows an accurate genotype call for a polymorphic SNP (in this case ¼ AA, ½ Aa, ¼ aa Mendelian segregation ratios, as expected for a biallelic codominant SNP segregating in a three-generation inbred F2 family). Panel B shows an accurate genotype call for a monomorphic SNP. Panel C shows a failed assay with low fluorescence intensity. The two grandparents of the F2 progeny are indicated by blue dots and the hybrid parent is indicated by green dots. The other colored dots correspond to the F2 offspring.
DownloadOther
Methodological specifications of the two approaches used to link genotypes and phenotypes.
DownloadArticle statistics
 Views: 14414
Views: 14414
Downloads
 PDF: 771
PDF: 771
